Cas9 SmartNuclease™ Extra Ligation Kit
Products
| Catalog Number | Description | Size | Price | Quantity | Add to Cart | |||
|---|---|---|---|---|---|---|---|---|
| CAS9LIG-KIT | Cas9 SmartNuclease Extra Ligation Kit (includes 5x ligation buffer (10 µL) and Fast ligase (2.5 µL)) | 1 Kit | $103 |
|
||||
Overview
Overview
Clone more gRNAs into our All-in-one Cas9 SmartNuclease Plasmids
In case you need more ligase and buffer than our All-in-one Cas9 SmartNuclease Kits supply, SBI offers additional Cas9 SmartNuclease™ 5x ligation buffer and Fast Ligase together as our Cas9 SmartNuclease Extra Ligation Kit.References
How It Works
How It Works
Using SBI’s All-in-one Cas9 SmartNuclease Plasmids
The workflow at-a-glance
- Design two DNA oligonucleotides that are sense and antisense sequences of the target DNA and are immediately upstream of a PAM sequence (5’ – NGG – 3’)
- Anneal the two oligonucleotides to generate a duplex
- Ligate the duplex into the pre-linearized All-in-one Cas9 SmartNuclease Plasmid
- Transform into competent cells and grow in LB/Kanamycin plate (50 µg/ml)
- Confirm positive clones by direct sequencing
- Transfect sequence-verified All-in-one construct into mammalian cells using standard transfection protocols (co-transfect with an HR Targeting Vector, if required for your application)
- Perform a Surveyor Nuclease assay (or other suitable mismatch cleavage assays) to check for site-specific genome cleavage and select for desired clones
Selecting Target DNA Sequences
The selection of the target DNA sequence is not limited by any constraints, with exception of the requirement of a PAM sequence in the form of 5’ – NGG – 3’ (where N = any base) immediately following the target sequence. The typical length of the target sequence is 20 bp.
Genome Engineering
For more general guidance on using CRISPR/Cas9 technology for genome engineering, take a look at our CRISPR/Cas9 tutorials as well as the following application notes:
CRISPR/Cas9 Gene Knock-Out Application Note (PDF) »
CRISPR/Cas9 Gene Editing Application Note (PDF) »
CRISPR/Cas9 Gene Tagging Application Note (PDF) »
CRISPR/Cas9 Basics
Through careful selection of the target sequence and design of a donor plasmid for homologous recombination, you can achieve efficient and highly targeted genomic modification with CRISPR/Cas9.
The system
Cas9 protein—uses guide RNA (gRNA) to direct site-specific, double-strand DNA cleavage adjacent to a protospacer adapter motif (PAM) in the target DNA.
gRNA—RNA sequence that guides Cas9 to cleave a homologous region in the target genome. Efficient cleavage only where the gRNA homology is adjacent to a PAM.
PAM—protospacer adapter motif, NGG, is a target DNA sequence that spCas9 will cut upstream from if directed to by the gRNA.
The workflow at-a-glance
DESIGN: Select gRNA and HR donor plasmids. Choice of gRNA site and design of donor plasmid determines whether the homologous recombination event results in a knock-out, knock-in, edit, or tagging.
CONSTRUCT: Clone gRNA into all-in-one Cas9 vector. Clone 5’ and 3’ homology arms into HR donor plasmid. If creating a knock-in, clone desired gene into HR donor.
CO-TRANSFECT or CO-INJECT: Introduce Cas9, gRNA, and HR Donors into the target cells using co-transfection for plasmids, co-transduction for lentivirus, or co-injection for mRNAs.
SELECT/SCREEN: Select or screen for mutants and verify.
VALIDATE: Genotype or sequence putative mutants to verify single or biallelic conversion.
Supporting Data
Supporting Data
Genome engineering with SBI’s All-in-one Cas9 SmartNuclease Plasmids
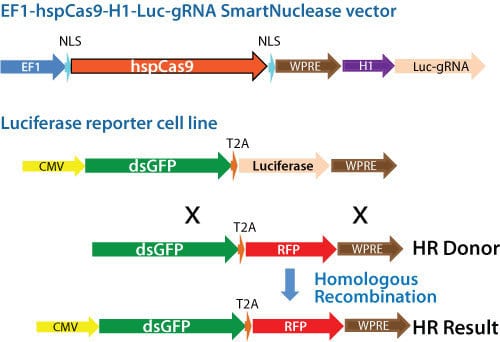
Figure 1. Schematic showing knock-out of a genomic luciferase gene by insertion of an RFP gene. Cas9 activity and gRNA were delivered using an All-in-one Cas9 SmartNuclease Plasmid.
To demonstrate the capabilities of our All-in-one Cas9 SmartNuclease Plasmids and also show how the choice of gRNA sequence can affect genome editing efficiencies, we designed a study to knock-out a genomic copy of a luciferase gene by inserting a copy of the RFP gene. This study used two different All-in-one constructs—EF1-hspCas9-H1-Luc-gRNA1 and EF1-hspCas9-H1-Luc-gRNA2—and an HR donor vector containing the RFP gene surrounded by sequences homologous to the DNA around the genomic luciferase gene (Figure 1). While both constructs enabled successful genome editing, as indicated by the recovery of RFP-positive clones, the construct with gRNA1 provided more efficient genome editing than the construct with gRNA2 (Figure 2).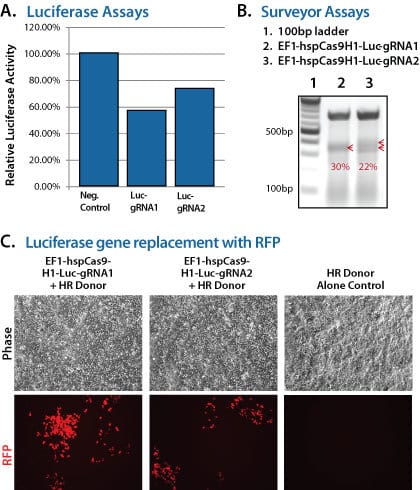
Figure 2. The All-in-one Cas9 SmartNuclease Plasmid enabled successful genomic luciferase knock-out and replacement with RFP. (A) Insertion of the RFP gene into the genomically-integrated luciferase gene results in a decrease in luciferase activity, indicating successful genome editing. (B) A Surveyor Nuclease Assay reflects finding from (A) that the gRNA1 construct delivered slightly more cleavage than the gRNA2 construct. (C) Fluorescence imaging shows successful genome editing by the recovery of RFP-positive clones.
FAQs
Resources
Citations
Related Products
Products
| Catalog Number | Description | Size | Price | Quantity | Add to Cart | |||
|---|---|---|---|---|---|---|---|---|
| CAS9LIG-KIT | Cas9 SmartNuclease Extra Ligation Kit (includes 5x ligation buffer (10 µL) and Fast ligase (2.5 µL)) | 1 Kit | $103 |
|
||||
Overview
Overview
Clone more gRNAs into our All-in-one Cas9 SmartNuclease Plasmids
In case you need more ligase and buffer than our All-in-one Cas9 SmartNuclease Kits supply, SBI offers additional Cas9 SmartNuclease™ 5x ligation buffer and Fast Ligase together as our Cas9 SmartNuclease Extra Ligation Kit.References
How It Works
How It Works
Using SBI’s All-in-one Cas9 SmartNuclease Plasmids
The workflow at-a-glance
- Design two DNA oligonucleotides that are sense and antisense sequences of the target DNA and are immediately upstream of a PAM sequence (5’ – NGG – 3’)
- Anneal the two oligonucleotides to generate a duplex
- Ligate the duplex into the pre-linearized All-in-one Cas9 SmartNuclease Plasmid
- Transform into competent cells and grow in LB/Kanamycin plate (50 µg/ml)
- Confirm positive clones by direct sequencing
- Transfect sequence-verified All-in-one construct into mammalian cells using standard transfection protocols (co-transfect with an HR Targeting Vector, if required for your application)
- Perform a Surveyor Nuclease assay (or other suitable mismatch cleavage assays) to check for site-specific genome cleavage and select for desired clones
Selecting Target DNA Sequences
The selection of the target DNA sequence is not limited by any constraints, with exception of the requirement of a PAM sequence in the form of 5’ – NGG – 3’ (where N = any base) immediately following the target sequence. The typical length of the target sequence is 20 bp.
Genome Engineering
For more general guidance on using CRISPR/Cas9 technology for genome engineering, take a look at our CRISPR/Cas9 tutorials as well as the following application notes:
CRISPR/Cas9 Gene Knock-Out Application Note (PDF) »
CRISPR/Cas9 Gene Editing Application Note (PDF) »
CRISPR/Cas9 Gene Tagging Application Note (PDF) »
CRISPR/Cas9 Basics
Through careful selection of the target sequence and design of a donor plasmid for homologous recombination, you can achieve efficient and highly targeted genomic modification with CRISPR/Cas9.
The system
Cas9 protein—uses guide RNA (gRNA) to direct site-specific, double-strand DNA cleavage adjacent to a protospacer adapter motif (PAM) in the target DNA.
gRNA—RNA sequence that guides Cas9 to cleave a homologous region in the target genome. Efficient cleavage only where the gRNA homology is adjacent to a PAM.
PAM—protospacer adapter motif, NGG, is a target DNA sequence that spCas9 will cut upstream from if directed to by the gRNA.
The workflow at-a-glance
DESIGN: Select gRNA and HR donor plasmids. Choice of gRNA site and design of donor plasmid determines whether the homologous recombination event results in a knock-out, knock-in, edit, or tagging.
CONSTRUCT: Clone gRNA into all-in-one Cas9 vector. Clone 5’ and 3’ homology arms into HR donor plasmid. If creating a knock-in, clone desired gene into HR donor.
CO-TRANSFECT or CO-INJECT: Introduce Cas9, gRNA, and HR Donors into the target cells using co-transfection for plasmids, co-transduction for lentivirus, or co-injection for mRNAs.
SELECT/SCREEN: Select or screen for mutants and verify.
VALIDATE: Genotype or sequence putative mutants to verify single or biallelic conversion.
Supporting Data
Supporting Data
Genome engineering with SBI’s All-in-one Cas9 SmartNuclease Plasmids
Figure 1. Schematic showing knock-out of a genomic luciferase gene by insertion of an RFP gene. Cas9 activity and gRNA were delivered using an All-in-one Cas9 SmartNuclease Plasmid.
To demonstrate the capabilities of our All-in-one Cas9 SmartNuclease Plasmids and also show how the choice of gRNA sequence can affect genome editing efficiencies, we designed a study to knock-out a genomic copy of a luciferase gene by inserting a copy of the RFP gene. This study used two different All-in-one constructs—EF1-hspCas9-H1-Luc-gRNA1 and EF1-hspCas9-H1-Luc-gRNA2—and an HR donor vector containing the RFP gene surrounded by sequences homologous to the DNA around the genomic luciferase gene (Figure 1). While both constructs enabled successful genome editing, as indicated by the recovery of RFP-positive clones, the construct with gRNA1 provided more efficient genome editing than the construct with gRNA2 (Figure 2).
Figure 2. The All-in-one Cas9 SmartNuclease Plasmid enabled successful genomic luciferase knock-out and replacement with RFP. (A) Insertion of the RFP gene into the genomically-integrated luciferase gene results in a decrease in luciferase activity, indicating successful genome editing. (B) A Surveyor Nuclease Assay reflects finding from (A) that the gRNA1 construct delivered slightly more cleavage than the gRNA2 construct. (C) Fluorescence imaging shows successful genome editing by the recovery of RFP-positive clones.