PrecisionX™ Multiplex gRNA Cloning Kit
- Edit multiple loci at once, save time and reagents
- Easily generate multi-cistronic gRNA expression
- constructs
- Ideal for Cas9 Nickase applications
- Great for studying signaling pathways
Products
| Catalog Number | Description | Size | Price | Quantity | Add to Cart | |||
|---|---|---|---|---|---|---|---|---|
| CAS9-GRNA-KIT | PrecisionX Multiplex gRNA Cloning Kit | 10 Reactions | $532 |
|
||||
Overview
Overview
Speeding your multi-gene CRISPR/Cas9 projects
With the PrecisionX™ Multiplex gRNA Cloning Kit, you can place multiple gRNAs into a single construct for faster, more efficient genome editing at multiple genomic locations. Compatible with any of SBI’s All-in-one Cas9 SmartNuclease Plasmids and gRNA cloning constructs as well as other popular Cas9/gRNA cloning vectors, such as pX330, pX335, pX458, and pX459, the Multiplex gRNA Cloning Kit is an easy-to-use tool for accelerating your CRISPR/Cas9 projects.
- Edit multiple loci at once, save time and reagents
- Easily generate multi-cistronic gRNA expression
- constructs
- Ideal for Cas9 Nickase applications
- Great for studying signaling pathways
References
How It Works
How It Works
Using the Multiplex gRNA Cloning Kit
Figure 1. The Multiplex gRNA Cloning Kit workflow.
The PrecisionX Multiplex gRNA Cloning Kit provides H1 and U6 promoter blocks to easily build multiple gRNA cassettes (Figure 1). In Step 1, primers with overlapping ends containing the desired gRNA (designed by user) and the scaffold-promoter block (from the kit) are combined in a PCR reaction to generate a PCR amplicon containing both gRNAs. In Step 2, the PCR amplicon is combined with the fusion reaction mix along with the linearized expression vector for seamless construction.
The cloning process is extremely efficient and the kit can be adapted to two or more gRNAs in a single fusion reaction.
Genome engineering with CRISPR/Cas9
For general guidance on using CRISPR/Cas9 technology for genome engineering, take a look at our CRISPR/Cas9 tutorials as well as the following application notes:
CRISPR/Cas9 Gene Knock-Out Application Note (PDF) »
CRISPR/Cas9 Gene Editing Application Note (PDF) »
CRISPR/Cas9 Gene Tagging Application Note (PDF) »
CRISPR/Cas9 Basics
Through careful selection of the target sequence and design of a donor plasmid for homologous recombination, you can achieve efficient and highly targeted genomic modification with CRISPR/Cas9.
Cas9 protein—uses guide RNA (gRNA) to direct site-specific, double-strand DNA cleavage adjacent to a protospacer adapter motif (PAM) in the target DNA.
gRNA—RNA sequence that guides Cas9 to cleave a homologous region in the target genome. Efficient cleavage only where the gRNA homology is adjacent to a PAM.
PAM—protospacer adapter motif, NGG, is a target DNA sequence that spCas9 will cut upstream from if directed to by the gRNA.
The workflow at-a-glance
DESIGN: Select gRNA and HR donor plasmids. Choice of gRNA site and design of donor plasmid determines whether the homologous recombination event results in a knock-out, knock-in, edit, or tagging.
CONSTRUCT: Clone gRNA into all-in-one Cas9 vector. Clone 5’ and 3’ homology arms into HR donor plasmid. If creating a knock-in, clone desired gene into HR donor.
CO-TRANSFECT or CO-INJECT: Introduce Cas9, gRNA, and HR Donors into the target cells using co-transfection for plasmids, co-transduction for lentivirus, or co-injection for mRNAs.
SELECT/SCREEN: Select or screen for mutants and verify.
VALIDATE: Genotype or sequence putative mutants to verify single or biallelic conversion.
Supporting Data
Supporting Data
Simultaneous editing at multiple genomic locations
Figure 2. The Multiplex gRNA Cloning Kit enables efficient delivery of four gRNAs.(A) We used a quad-plex gRNA created using the Multilplex gRNA Cloning Kit and an All-in-one Cas9 SmartNickase Vector to remove a (B) 1260 bp GFP-T2A-RFP segment from a cell line with a stably integrated CMV-GFP-T2A-RFP expression cassette. We cloned the four gRNAs into a Cas9 SmartNickase vector (EF1Nickase-H1-gRNA) to guide two double nicking events—one at the 5’ end of the GFP and the other at the 3’ end of the RFP gene. (C) PCR assays with primers just outside of the GFP and RFP genes generate a 1600 bp fragment in the absence of the SmartNickase vector (lane 1), and a 340 bp fragment in the presence of the Cas9 SmartNickase-4 gRNA construct (lane 2), demonstrating the efficiency of SmartNickase-mediated paired double-nicking and GFP-T2A-RFP genomic deletion. (D) Deletion of both GFP and RFP activities can also be seen in a functional assay, through reduction in both GFP and RFP fluorescence.
FAQs
Documentation
Citations
Related Products
Products
| Catalog Number | Description | Size | Price | Quantity | Add to Cart | |||
|---|---|---|---|---|---|---|---|---|
| CAS9-GRNA-KIT | PrecisionX Multiplex gRNA Cloning Kit | 10 Reactions | $532 |
|
||||
Overview
Overview
Speeding your multi-gene CRISPR/Cas9 projects
With the PrecisionX™ Multiplex gRNA Cloning Kit, you can place multiple gRNAs into a single construct for faster, more efficient genome editing at multiple genomic locations. Compatible with any of SBI’s All-in-one Cas9 SmartNuclease Plasmids and gRNA cloning constructs as well as other popular Cas9/gRNA cloning vectors, such as pX330, pX335, pX458, and pX459, the Multiplex gRNA Cloning Kit is an easy-to-use tool for accelerating your CRISPR/Cas9 projects.
- Edit multiple loci at once, save time and reagents
- Easily generate multi-cistronic gRNA expression
- constructs
- Ideal for Cas9 Nickase applications
- Great for studying signaling pathways
References
How It Works
How It Works
Using the Multiplex gRNA Cloning Kit
Figure 1. The Multiplex gRNA Cloning Kit workflow.
The PrecisionX Multiplex gRNA Cloning Kit provides H1 and U6 promoter blocks to easily build multiple gRNA cassettes (Figure 1). In Step 1, primers with overlapping ends containing the desired gRNA (designed by user) and the scaffold-promoter block (from the kit) are combined in a PCR reaction to generate a PCR amplicon containing both gRNAs. In Step 2, the PCR amplicon is combined with the fusion reaction mix along with the linearized expression vector for seamless construction.
The cloning process is extremely efficient and the kit can be adapted to two or more gRNAs in a single fusion reaction.
Genome engineering with CRISPR/Cas9
For general guidance on using CRISPR/Cas9 technology for genome engineering, take a look at our CRISPR/Cas9 tutorials as well as the following application notes:
CRISPR/Cas9 Gene Knock-Out Application Note (PDF) »
CRISPR/Cas9 Gene Editing Application Note (PDF) »
CRISPR/Cas9 Gene Tagging Application Note (PDF) »
CRISPR/Cas9 Basics
Through careful selection of the target sequence and design of a donor plasmid for homologous recombination, you can achieve efficient and highly targeted genomic modification with CRISPR/Cas9.
Cas9 protein—uses guide RNA (gRNA) to direct site-specific, double-strand DNA cleavage adjacent to a protospacer adapter motif (PAM) in the target DNA.
gRNA—RNA sequence that guides Cas9 to cleave a homologous region in the target genome. Efficient cleavage only where the gRNA homology is adjacent to a PAM.
PAM—protospacer adapter motif, NGG, is a target DNA sequence that spCas9 will cut upstream from if directed to by the gRNA.
The workflow at-a-glance
DESIGN: Select gRNA and HR donor plasmids. Choice of gRNA site and design of donor plasmid determines whether the homologous recombination event results in a knock-out, knock-in, edit, or tagging.
CONSTRUCT: Clone gRNA into all-in-one Cas9 vector. Clone 5’ and 3’ homology arms into HR donor plasmid. If creating a knock-in, clone desired gene into HR donor.
CO-TRANSFECT or CO-INJECT: Introduce Cas9, gRNA, and HR Donors into the target cells using co-transfection for plasmids, co-transduction for lentivirus, or co-injection for mRNAs.
SELECT/SCREEN: Select or screen for mutants and verify.
VALIDATE: Genotype or sequence putative mutants to verify single or biallelic conversion.
Supporting Data
Supporting Data
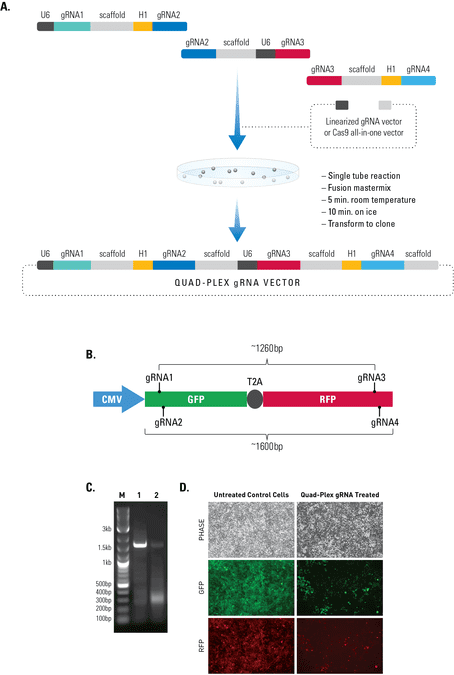
Simultaneous editing at multiple genomic locations
Figure 2. The Multiplex gRNA Cloning Kit enables efficient delivery of four gRNAs.(A) We used a quad-plex gRNA created using the Multilplex gRNA Cloning Kit and an All-in-one Cas9 SmartNickase Vector to remove a (B) 1260 bp GFP-T2A-RFP segment from a cell line with a stably integrated CMV-GFP-T2A-RFP expression cassette. We cloned the four gRNAs into a Cas9 SmartNickase vector (EF1Nickase-H1-gRNA) to guide two double nicking events—one at the 5’ end of the GFP and the other at the 3’ end of the RFP gene. (C) PCR assays with primers just outside of the GFP and RFP genes generate a 1600 bp fragment in the absence of the SmartNickase vector (lane 1), and a 340 bp fragment in the presence of the Cas9 SmartNickase-4 gRNA construct (lane 2), demonstrating the efficiency of SmartNickase-mediated paired double-nicking and GFP-T2A-RFP genomic deletion. (D) Deletion of both GFP and RFP activities can also be seen in a functional assay, through reduction in both GFP and RFP fluorescence.