pGreenFire 2.0 NFκB Reporter Lentivector & Virus
- Sort responsive cells with GFP
- Measure activity with red firefly luciferase
- Leverage SBI’s highly-regarded lentivectors
- Create stable signaling pathway reporter cell lines
- Introduce reporters into difficult-to-transfect cell types, including primary and non-dividing mammalian cell lines
Products
| Catalog Number | Description | Size | Price | Quantity | Add to Cart | |||
|---|---|---|---|---|---|---|---|---|
| TR412PA-P | pGreenFire 2.0 NFkB reporter plasmid (pGF2-NFκB-rFluc-T2A-GFP-mPGK-Puro) | 10 µg | $941 |
|
||||
| TR412VA-P | pGreenFire 2.0 NFkB reporter virus (pGF2-NFκB-rFluc-T2A-GFP-mPGK-Puro) | >2 x 10^6 IFUs | $941 |
|
||||
Overview
Overview
Monitor signal transduction in real time with our re-engineered pGreenFire 2.0 Lentivectors
SBI has upgraded our popular pGreenFire signaling pathway reporter lentivectors with a design that leads to more reliable generation of stable cell lines. We’ve also swapped in the red firefly luciferase reporter (rFLuc), which opens up the possibility of performing a dual-spectral luciferase assay and delivers greater sensitivity for in vivo applications than conventional luciferase.
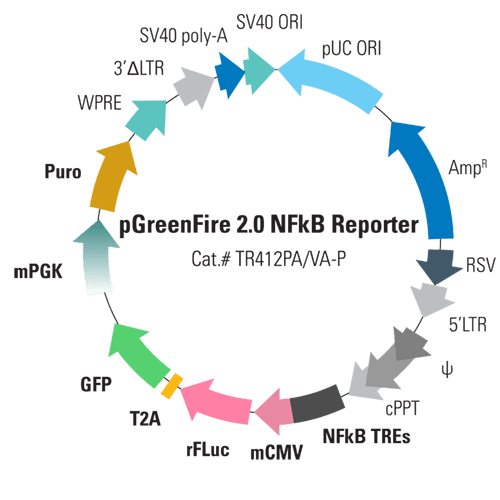
With the pGreenFire 2.0 NFκB Reporter Lentivector & Virus (pGF2-NFκB-rFluc-T2A-GFP-mPGK-Puro), the core reporter functionality is similar to the original pGreenFire lentivector—NF-κB transcriptional response elements (TREs) are placed upstream of a minimal CMV promoter (mCMV) which together drive co-expression of rFLuc and GFP in response to NF-κB activity. The result is the ability to quantitatively measure NF-κB activity using both fluorescence and luciferase activity.
What makes our next-gen pGreenFire 2.0 vectors even better than other TRE reporter vectors is the smart design, which adds in a constitutive selection cassette for stable cell line generation while minimizing interference with the upstream TRE. By using a weak/moderate mPGK promoter to drive the antibiotic selection marker (puromycin resistance) and carefully arranging the conditional reporter genes, the selection marker is reliably expressed without compromising conditional expression of rFLuc and GFP.
As with our original pGreenFire1 vectors, all pGreenFire 2.0 lentivectors leverage our reliable lentivector technology and save you time with pre-built signal transduction pathway reporters that come as ready-to-transduce pre-packaged lentivirus and plasmid that can be transfected into the lentivirus producing system of your choice*.
- Sort responsive cells with GFP
- Measure activity with red firefly luciferase
- Leverage SBI’s highly-regarded lentivectors
- Create stable signaling pathway reporter cell lines
- Introduce reporters into difficult-to-transfect cell types, including primary and non-dividing mammalian cell lines
 *Please note that these vectors only function properly when transduced. Transfection keeps the constitutive RSV promoter intact, leading to nonspecific expression of the reporter genes.
*Please note that these vectors only function properly when transduced. Transfection keeps the constitutive RSV promoter intact, leading to nonspecific expression of the reporter genes.References
How It Works
Supporting Data
Supporting Data
See our pGreenFire 2.0 transcriptional response element reporters in action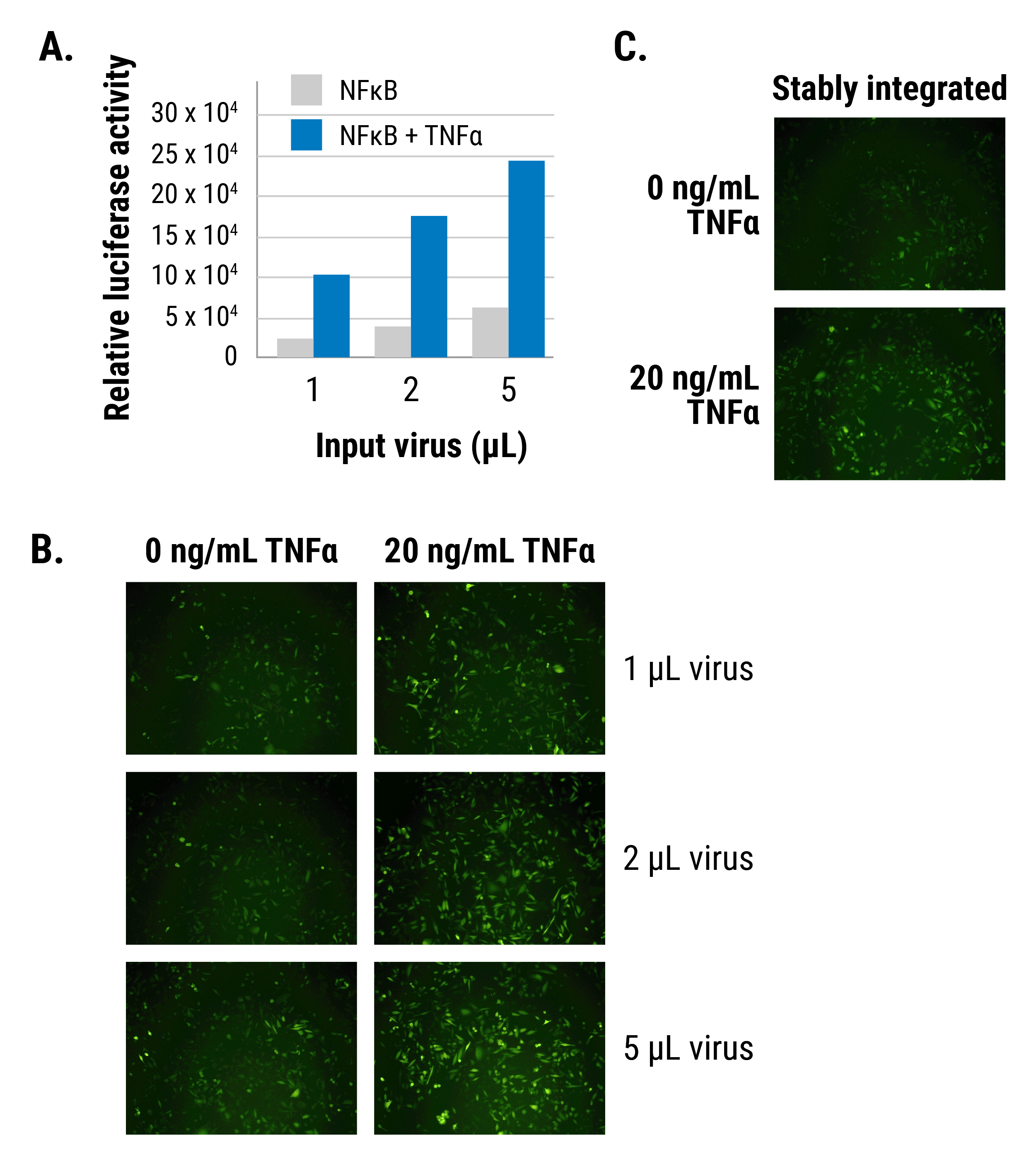
Figure 1.The pGreenFire 2.0 NFκB Reporter efficiently and quantitatively reports on NFκB activity in MDA-MB-213 cells. Relative luciferase activity (A) and GFP activity (B) both increase in response to TNFα, and NFκB inducer. (C) Like all pGreenFire 2.0 lentivectors, the pGreenFire 2.0 NFκB Reporter contains an mPGK-Puro cassette to streamline creation of stable reporters integrated into the cell lines of your choice.
FAQs
Documentation
Citations
Related Products
Products
| Catalog Number | Description | Size | Price | Quantity | Add to Cart | |||
|---|---|---|---|---|---|---|---|---|
| TR412PA-P | pGreenFire 2.0 NFkB reporter plasmid (pGF2-NFκB-rFluc-T2A-GFP-mPGK-Puro) | 10 µg | $941 |
|
||||
| TR412VA-P | pGreenFire 2.0 NFkB reporter virus (pGF2-NFκB-rFluc-T2A-GFP-mPGK-Puro) | >2 x 10^6 IFUs | $941 |
|
||||
Overview
Overview
Monitor signal transduction in real time with our re-engineered pGreenFire 2.0 Lentivectors
SBI has upgraded our popular pGreenFire signaling pathway reporter lentivectors with a design that leads to more reliable generation of stable cell lines. We’ve also swapped in the red firefly luciferase reporter (rFLuc), which opens up the possibility of performing a dual-spectral luciferase assay and delivers greater sensitivity for in vivo applications than conventional luciferase.
With the pGreenFire 2.0 NFκB Reporter Lentivector & Virus (pGF2-NFκB-rFluc-T2A-GFP-mPGK-Puro), the core reporter functionality is similar to the original pGreenFire lentivector—NF-κB transcriptional response elements (TREs) are placed upstream of a minimal CMV promoter (mCMV) which together drive co-expression of rFLuc and GFP in response to NF-κB activity. The result is the ability to quantitatively measure NF-κB activity using both fluorescence and luciferase activity.
What makes our next-gen pGreenFire 2.0 vectors even better than other TRE reporter vectors is the smart design, which adds in a constitutive selection cassette for stable cell line generation while minimizing interference with the upstream TRE. By using a weak/moderate mPGK promoter to drive the antibiotic selection marker (puromycin resistance) and carefully arranging the conditional reporter genes, the selection marker is reliably expressed without compromising conditional expression of rFLuc and GFP.
As with our original pGreenFire1 vectors, all pGreenFire 2.0 lentivectors leverage our reliable lentivector technology and save you time with pre-built signal transduction pathway reporters that come as ready-to-transduce pre-packaged lentivirus and plasmid that can be transfected into the lentivirus producing system of your choice*.
- Sort responsive cells with GFP
- Measure activity with red firefly luciferase
- Leverage SBI’s highly-regarded lentivectors
- Create stable signaling pathway reporter cell lines
- Introduce reporters into difficult-to-transfect cell types, including primary and non-dividing mammalian cell lines
 *Please note that these vectors only function properly when transduced. Transfection keeps the constitutive RSV promoter intact, leading to nonspecific expression of the reporter genes.
*Please note that these vectors only function properly when transduced. Transfection keeps the constitutive RSV promoter intact, leading to nonspecific expression of the reporter genes.References
How It Works
Supporting Data
Supporting Data
See our pGreenFire 2.0 transcriptional response element reporters in action
Figure 1.The pGreenFire 2.0 NFκB Reporter efficiently and quantitatively reports on NFκB activity in MDA-MB-213 cells. Relative luciferase activity (A) and GFP activity (B) both increase in response to TNFα, and NFκB inducer. (C) Like all pGreenFire 2.0 lentivectors, the pGreenFire 2.0 NFκB Reporter contains an mPGK-Puro cassette to streamline creation of stable reporters integrated into the cell lines of your choice.

