QuantiMir™ miR-16 Reference Assays
- Ready-to-use reference assay
- Designed for use with the QuantiMir Kit
- Suitable for high-throughput screening of clinical samples
Products
| Catalog Number | Description | Size | Price | Quantity | Add to Cart | |||
|---|---|---|---|---|---|---|---|---|
| RA420A-miR-16 | QuantiMir™ miR-16 Reference Assays | 50 Assays | $135 |
|
||||
Overview
Overview
Supporting your studies with ready-to-go reference assays
QuantiMir™ is SBI's simple and robust technology for profiling miRNAs. For labs using our QuantiMir Kit who need additional miR-16 assays for human or mouse, we offer this stand-alone set of 50 assays.
Visit the QuantiMir page to order the complete kit.
See all of our reference assays:| Catalog # | Amplified small RNA | Species compatibility |
|---|---|---|
| RA420A-hU6 | Human U6 | Human |
| RA420A-miR-16 | miR-16 | Human, Mouse |
| RA420A-mU6 | Mouse U6 | Mouse |
| RA420A-RNU1 | RNU1 (U1) | Human, Mouse, Rat, Monkey |
| RA420A-RNU43 | RNU43 | Human, Mouse, Rat |
| RA420A-rU6 | Rat U6 | Rat |
References
How It Works
How the QuantiMir Kit Works
The QuantiMir Kit enables robust miRNA quantitation through a simple and efficient workflow
Highly efficient poly-A tailing and reverse transcription in a single reaction tube provides uniform cDNA synthesis of miRNAs. The optimized reaction conditions and buffer components maximize cDNA yield when starting with several micrograms down to picograms of input total RNA. The universal 3′ tag sequence incorporated during reverse transcription enables easily scalable and accurate miRNA expression analysis by qPCR—profile thousands of different miRNAs from a single reverse transcription reaction.
- Tag all small RNAs with a poly-A tail
- Anneal an oligo-dT adaptor
- Reverse transcribe to create first-strand cDNA
The result is a cDNA pool of anchor-tailed miRNAs that are ready for qPCR.
Create custom miRNA assays
To design your own miRNA assays, simply synthesize an oligo using the sequence of the mature miRNA you’d like to profile as the forward primer in your miRNA qPCR assay, and use with the universal reverse primer included in the QuantiMir Kit.
Supporting Data
Supporting Data for the QuantiMir Kit
Converting miRNA into cDNA for accurate qPCR profiling and quantitation
The QuantiMir Kit is sensitive
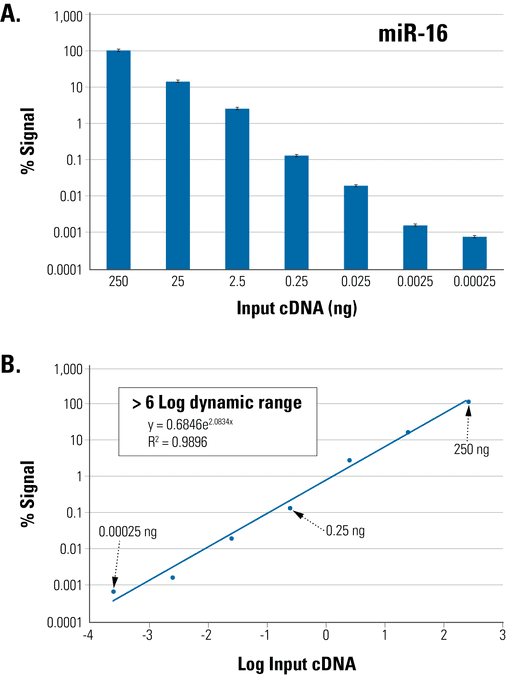
Figure 1. The QuantiMir Kit is highly sensitive and enables measurement across a wide dynamic range. (A) Starting with total, Trizol-extracted RNA or fractionated small RNA samples you can measure from several micrograms down to picograms of input RNA with excellent accuracy. (B) You can also detect differential miRNA expression across a dynamic range of at least 6 logs.
The QuantiMir Kit is accurate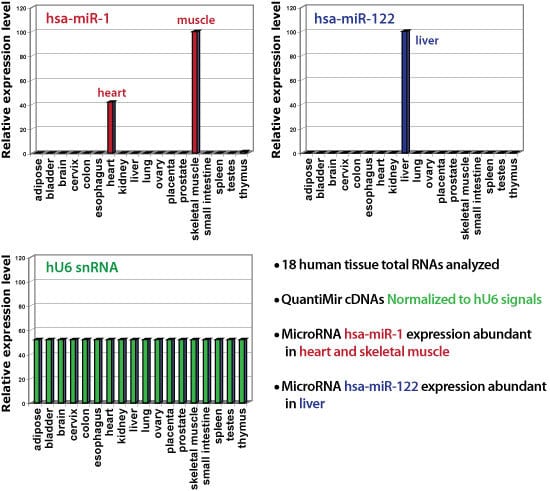
Figure 2. The QuantiMir Kit is accurate. We used the QuantiMir Kit to synthesize first strand cDNAs from 18 different human tissues: adipose, bladder, brain, cervix, colon, esophagus, heart, kidney, liver, lung, ovary, placenta, prostate, skeletal muscle, small intestine, spleen, testes, and thymus. The cDNAs were balanced to yield equal Ct values for the U6 snRNA normalizing transcript (bottom left plot, green bars). Real-time PCR results demonstrate that the normalizing snRNA is uniformly expressed across the 18 tissues examined. As expected, assays for miR-1 are show specific expression in heart and musculoskeletal tissues (top left plot, red bars) whereas assays for miR-122 show specific expression in the liver (top right plot, blue bar).
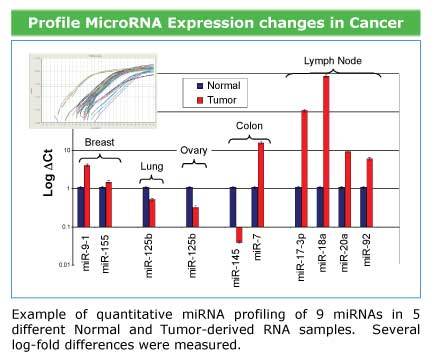
Use the QuantiMir Kit to profile miRNAs from cancerous and normal tissuesFigure 3. Use the QuantiMir Kit to profile miRNAs from cancerous and normal tissues. Example of quantitative miRNA profiling of nine miRNAs in five different normal and tumor-derived samples.
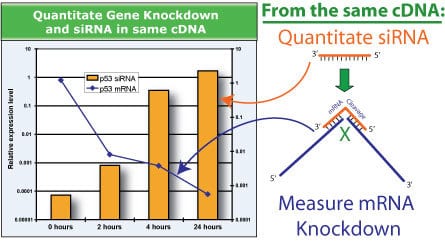
Measure both siRNA and mRNA knockdown in a single QuantiMir cDNA sampleFigure 4. Measure both siRNA and mRNA knockdown in a single QuantiMir cDNA sample. A time-course using anti-p53 shRNA-directed knockdown of the endogenous p53 mRNA transcript demonstrates how the QuantiMir Kit can be used to measure both p53 siRNA (orange bars) and p53 mRNA (blue line) via qPCR.
FAQs
Documentation
Citations
Related Products
Products
| Catalog Number | Description | Size | Price | Quantity | Add to Cart | |||
|---|---|---|---|---|---|---|---|---|
| RA420A-miR-16 | QuantiMir™ miR-16 Reference Assays | 50 Assays | $135 |
|
||||
Overview
Overview
Supporting your studies with ready-to-go reference assays
QuantiMir™ is SBI's simple and robust technology for profiling miRNAs. For labs using our QuantiMir Kit who need additional miR-16 assays for human or mouse, we offer this stand-alone set of 50 assays.
Visit the QuantiMir page to order the complete kit.
See all of our reference assays:| Catalog # | Amplified small RNA | Species compatibility |
|---|---|---|
| RA420A-hU6 | Human U6 | Human |
| RA420A-miR-16 | miR-16 | Human, Mouse |
| RA420A-mU6 | Mouse U6 | Mouse |
| RA420A-RNU1 | RNU1 (U1) | Human, Mouse, Rat, Monkey |
| RA420A-RNU43 | RNU43 | Human, Mouse, Rat |
| RA420A-rU6 | Rat U6 | Rat |
References
How It Works
How the QuantiMir Kit Works
The QuantiMir Kit enables robust miRNA quantitation through a simple and efficient workflow
Highly efficient poly-A tailing and reverse transcription in a single reaction tube provides uniform cDNA synthesis of miRNAs. The optimized reaction conditions and buffer components maximize cDNA yield when starting with several micrograms down to picograms of input total RNA. The universal 3′ tag sequence incorporated during reverse transcription enables easily scalable and accurate miRNA expression analysis by qPCR—profile thousands of different miRNAs from a single reverse transcription reaction.
- Tag all small RNAs with a poly-A tail
- Anneal an oligo-dT adaptor
- Reverse transcribe to create first-strand cDNA
The result is a cDNA pool of anchor-tailed miRNAs that are ready for qPCR.
Create custom miRNA assays
To design your own miRNA assays, simply synthesize an oligo using the sequence of the mature miRNA you’d like to profile as the forward primer in your miRNA qPCR assay, and use with the universal reverse primer included in the QuantiMir Kit.
Supporting Data
Supporting Data for the QuantiMir Kit
Converting miRNA into cDNA for accurate qPCR profiling and quantitation
The QuantiMir Kit is sensitive
Figure 1. The QuantiMir Kit is highly sensitive and enables measurement across a wide dynamic range. (A) Starting with total, Trizol-extracted RNA or fractionated small RNA samples you can measure from several micrograms down to picograms of input RNA with excellent accuracy. (B) You can also detect differential miRNA expression across a dynamic range of at least 6 logs.
The QuantiMir Kit is accurate
Figure 2. The QuantiMir Kit is accurate. We used the QuantiMir Kit to synthesize first strand cDNAs from 18 different human tissues: adipose, bladder, brain, cervix, colon, esophagus, heart, kidney, liver, lung, ovary, placenta, prostate, skeletal muscle, small intestine, spleen, testes, and thymus. The cDNAs were balanced to yield equal Ct values for the U6 snRNA normalizing transcript (bottom left plot, green bars). Real-time PCR results demonstrate that the normalizing snRNA is uniformly expressed across the 18 tissues examined. As expected, assays for miR-1 are show specific expression in heart and musculoskeletal tissues (top left plot, red bars) whereas assays for miR-122 show specific expression in the liver (top right plot, blue bar).
Use the QuantiMir Kit to profile miRNAs from cancerous and normal tissuesFigure 3. Use the QuantiMir Kit to profile miRNAs from cancerous and normal tissues. Example of quantitative miRNA profiling of nine miRNAs in five different normal and tumor-derived samples.
Measure both siRNA and mRNA knockdown in a single QuantiMir cDNA sampleFigure 4. Measure both siRNA and mRNA knockdown in a single QuantiMir cDNA sample. A time-course using anti-p53 shRNA-directed knockdown of the endogenous p53 mRNA transcript demonstrates how the QuantiMir Kit can be used to measure both p53 siRNA (orange bars) and p53 mRNA (blue line) via qPCR.